tree-sitter-fasta
tree-sitter-fasta is a parser for the FASTA file format, commonly used for genomics and computational biology.
It based on Tree-sitter, which is commonly supported by many text editors.
tree-sitter-fasta can be used both as a language grammar in a text editor or as a parser in a programming language supported by Tree-sitter's language bindings.
Features
- Supports both nucleic and amino acid FASTA files
- Syntax highlighting
- Colours for all nucleobases
- Colours for methionine and stop codons
- Reference name and
:-separated highlights in description lines
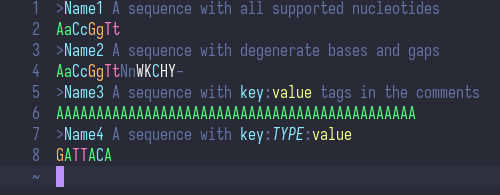
Syntax highlighting for a nucleic acid FASTA file.
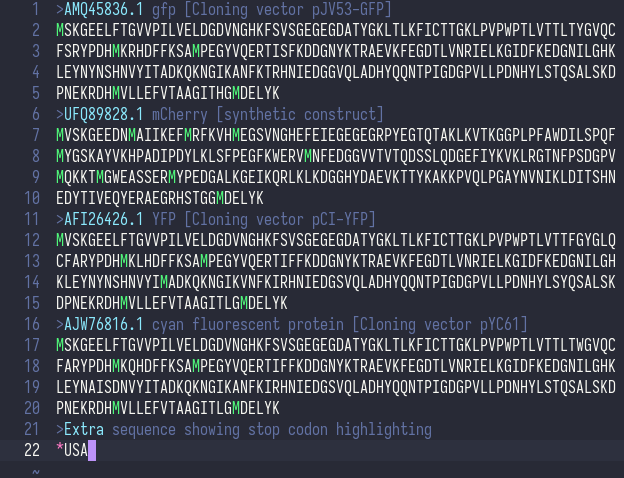
Syntax highlighting for an amino acid FASTA file.
Get started
Install the grammar following your text editor's instructions or install the package for a supported programming language with Tree-sitter bindings. See Installation for details.
Related projects
These other projects have no affiliation with this one, but may be of interest:
tree-sitter-fasta: An alternative Tree-sitter grammar for FASTA files by Will Gebbiefasta.el: An Emacs package that defines a major mode for FASTA files by Dave Pearson- sequed: DNA sequence editor and alignment viewer for emacs by Bruce Rannala
- Vim syntax for FASTA: A Vim syntax that detects FASTA files automatically
bioSyntax: Syntax highlighting for computational biology in various text editors- FASTA Sequence Highlighter: FASTA file formatting for Visual Studio Code